Rats: A Normal Hierarchical Model¶
An example from OpenBUGS [44] and section 6 of Gelfand et al. [29] concerning 30 rats whose weights were measured at each of five consecutive weeks.
Model¶
Weights are modeled as

where  is repeated weight measurement
is repeated weight measurement 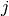 on rat
on rat 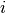 , and
, and  is the day on which the measurement was taken.
is the day on which the measurement was taken.
Analysis Program¶
using Mamba
## Data
rats = Dict{Symbol, Any}(
:y =>
[151, 199, 246, 283, 320,
145, 199, 249, 293, 354,
147, 214, 263, 312, 328,
155, 200, 237, 272, 297,
135, 188, 230, 280, 323,
159, 210, 252, 298, 331,
141, 189, 231, 275, 305,
159, 201, 248, 297, 338,
177, 236, 285, 350, 376,
134, 182, 220, 260, 296,
160, 208, 261, 313, 352,
143, 188, 220, 273, 314,
154, 200, 244, 289, 325,
171, 221, 270, 326, 358,
163, 216, 242, 281, 312,
160, 207, 248, 288, 324,
142, 187, 234, 280, 316,
156, 203, 243, 283, 317,
157, 212, 259, 307, 336,
152, 203, 246, 286, 321,
154, 205, 253, 298, 334,
139, 190, 225, 267, 302,
146, 191, 229, 272, 302,
157, 211, 250, 285, 323,
132, 185, 237, 286, 331,
160, 207, 257, 303, 345,
169, 216, 261, 295, 333,
157, 205, 248, 289, 316,
137, 180, 219, 258, 291,
153, 200, 244, 286, 324],
:x => [8.0, 15.0, 22.0, 29.0, 36.0]
)
rats[:xbar] = mean(rats[:x])
rats[:N] = size(rats[:y], 1)
rats[:T] = size(rats[:y], 2)
rats[:rat] = Int[div(i - 1, 5) + 1 for i in 1:150]
rats[:week] = Int[(i - 1) % 5 + 1 for i in 1:150]
rats[:X] = rats[:x][rats[:week]]
rats[:Xm] = rats[:X] - rats[:xbar]
## Model Specification
model = Model(
y = Stochastic(1,
(alpha, beta, rat, Xm, s2_c) ->
begin
mu = alpha[rat] + beta[rat] .* Xm
MvNormal(mu, sqrt(s2_c))
end,
false
),
alpha = Stochastic(1,
(mu_alpha, s2_alpha) -> Normal(mu_alpha, sqrt(s2_alpha)),
false
),
alpha0 = Logical(
(mu_alpha, xbar, mu_beta) -> mu_alpha - xbar * mu_beta
),
mu_alpha = Stochastic(
() -> Normal(0.0, 1000),
false
),
s2_alpha = Stochastic(
() -> InverseGamma(0.001, 0.001),
false
),
beta = Stochastic(1,
(mu_beta, s2_beta) -> Normal(mu_beta, sqrt(s2_beta)),
false
),
mu_beta = Stochastic(
() -> Normal(0.0, 1000)
),
s2_beta = Stochastic(
() -> InverseGamma(0.001, 0.001),
false
),
s2_c = Stochastic(
() -> InverseGamma(0.001, 0.001)
)
)
## Initial Values
inits = [
Dict(:y => rats[:y], :alpha => fill(250, 30), :beta => fill(6, 30),
:mu_alpha => 150, :mu_beta => 10, :s2_c => 1, :s2_alpha => 1,
:s2_beta => 1),
Dict(:y => rats[:y], :alpha => fill(20, 30), :beta => fill(0.6, 30),
:mu_alpha => 15, :mu_beta => 1, :s2_c => 10, :s2_alpha => 10,
:s2_beta => 10)
]
## Sampling Scheme
scheme = [Slice(:s2_c, 10.0),
AMWG(:alpha, 100.0),
Slice([:mu_alpha, :s2_alpha], [100.0, 10.0], Univariate),
AMWG(:beta, 1.0),
Slice([:mu_beta, :s2_beta], 1.0, Univariate)]
setsamplers!(model, scheme)
## MCMC Simulations
sim = mcmc(model, rats, inits, 10000, burnin=2500, thin=2, chains=2)
describe(sim)
Results¶
Iterations = 2502:10000
Thinning interval = 2
Chains = 1,2
Samples per chain = 3750
Empirical Posterior Estimates:
Mean SD Naive SE MCSE ESS
s2_c 37.2543133 6.026634572 0.0695895819 0.2337982327 664.4576
mu_beta 6.1830663 0.108042927 0.0012475723 0.0017921615 3634.4474
alpha0 106.6259925 3.459210115 0.0399435178 0.0526804390 3750.0000
Quantiles:
2.5% 25.0% 50.0% 75.0% 97.5%
s2_c 27.778388 33.0906026 36.4630047 40.5538472 51.5713716
mu_beta 5.969850 6.1110307 6.1836454 6.2538831 6.3964953
alpha0 99.815707 104.3369878 106.6105679 108.9124224 113.5045347