Dependent¶
AbstractDependent is an abstract type designed to store values and attributes of model nodes, including parameters 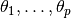 to be simulated via MCMC, functions of the parameters, and likelihood specifications on observed data. It extends the base
to be simulated via MCMC, functions of the parameters, and likelihood specifications on observed data. It extends the base Variate types with method functions defined for the fields summarized below. Like the type it extends, values are stored in a value field and can be used with method functions that accept Float64 or Array{Float64, N} type objects.
Since parameter values in the AbstractDependent structure are stored as a scalar or array, objects of this type can be created for model parameters of corresponding dimensions, with the choice between the two being user and application-specific. At one end of the spectrum, a model might be formulated in terms of parameters that are all scalars, with a separate instances of AbstractDependent for each one. At the other end, a formulation might be made in terms of a single parameter array, with one corresponding instance of AbstractDependent. Whether to formulate parameters as scalars or arrays will depend on the application at hand. Array formulations should be considered for parameters and data that have multivariate distributions, or are to be used as such in numeric operations and functions. In other cases, scalar parametrizations may be preferable. Situations in which parameter arrays are often used include the specification of regression coefficients and random effects.
Declaration¶
typealias AbstractDependent Union{AbstractLogical, AbstractStochastic}
Fields¶
value::T: scalar or array ofFloat64values that represent samples from a target distribution.symbol::Symbol: identifying symbol for the node.monitor::Vector{Int}: indices identifying elements of thevaluefield to include in monitored MCMC sampler output.eval::Function: function for updating the state of the node.sources::Vector{Symbol}: other nodes upon whom the values of this one depends.targets::Vector{Symbol}:Dependentnodes that depend on this one. Elements oftargetsare topologically sorted so that a given node in the vector is conditionally independent of subsequent nodes, given the previous ones.
Display¶
-
show(d::AbstractDependent)¶ Write a text representation of nodal values and attributes to the current output stream.
-
showall(d::AbstractDependent)¶ Write a verbose text representation of nodal values and attributes to the current output stream.
Initialization¶
-
setmonitor!(d::AbstractDependent, monitor::Bool)¶ -
setmonitor!(d::AbstractDependent, monitor::Vector{Int}) Specify node elements to be included in monitored MCMC sampler output.
Arguments
d: node whose elements contain sampled MCMC values.monitor: boolean indicating whether all elements are monitored, or vector of element-wise indices of elements to monitor.
Value
Returnsdwith itsmonitorfield updated to reflect the specified monitoring.
Node Operations¶
-
logpdf(d::AbstractDependent, transform::Bool=false)¶ -
logpdf(d::AbstractDependent, x, transform::Bool=false) Evaluate the log-density function for a node. In this method, no density function is assumed for the node, and a constant value of 0 is returned. This method function may be redefined for subtypes of
AbstractDependentthat have distributional specifications.Arguments
d: node for which to evaluate the log-density.x: value, of the same type and shape as the node value, at which to perform the evaluation. If not specified, the node value is used.transform: whether the evaluation is on the link-transformed scale.
Value
The resulting numeric value of the log-density.
-
unlist(d::AbstractDependent, transform::Bool=false)¶ -
unlist(d::AbstractDependent, x::Real, transform::Bool=false) -
unlist(d::AbstractDependent, x::AbstractArray, transform::Bool=false) -
relist(d::AbstractDependent, x::AbstractArray, transform::Bool=false)¶ Extract (unlist) node values to a vector, or re-assemble (relist) values to be put into a node. In this generic method, all values are listed. The methods are used internally for the extraction of unique stochastic node values to sample, and can be redefined to implement different behaviors for
AbstractDependentsubtypes.Arguments
d: node for which to unlist or relist values.x: values to be listed. If not specified, the node values are used.transform: whether to apply a link or inverse-link transformation to the values. In this generic method, transformations are defined to be the identity function.
Value
Returns unmodifiedxvalues as a vector (unlist) or in the same shape as the specified node (relist).
Logical¶
The Logical types inherit fields and method functions from the AbstractDependent type, and adds the constructors and methods listed below. It is designed for nodes that are deterministic functions of model parameters and data.
Declarations¶
type ScalarLogical <: ScalarVariate
type ArrayLogical{N} <: ArrayVariate{N}
typealias AbstractLogical Union{ScalarLogical, ArrayLogical}
Fields¶
value: values of typeFloat64forScalarLogicalnodes andArray{Float64}forArrayLogicalnodes that represent samples from a target distribution.symbol::Symbol: identifying symbol for the node.monitor::Vector{Int}: indices identifying elements of thevaluefield to include in monitored MCMC sampler output.eval::Function: function for updating values stored invalue.sources::Vector{Symbol}: other nodes upon whom the values of this one depends.targets::Vector{Symbol}:Dependentnodes that depend on this one. Elements oftargetsare topologically sorted so that a given node in the vector is conditionally independent of subsequent nodes, given the previous ones.
Constructors¶
-
Logical(f::Function, monitor::Union{Bool, Vector{Int}}=true)¶ -
Logical(d::Integer, f::Function, monitor::Union{Bool, Vector{Int}}=true) Construct a
Logicalobject that defines a logical model node.Arguments
d: number of dimensions for array nodes.f: function whose untyped arguments are the other model nodes upon which this one depends. The function may contain any valid julia expression or code block. It will be saved in theevalfield of the constructed logical node and should return a value in the same type as and with which to update the node’svaluefield.monitor: boolean indicating whether all elements are monitored, or vector of element-wise indices of elements to monitor.
Value
Returns anArrayLogicalif the dimension argumentdis specified, and aScalarLogicalif not.Example
See the Model Specification section of the tutorial.
Stochastic¶
The Stochastic types inherit fields and method functions from the AbstractDependent type, and adds the additional ones listed below. It is designed for model parameters or data that have distributional or likelihood specifications, respectively. Its stochastic relationship to other nodes and data structures is represented by the structure stored in distr field.
Declarations¶
type ScalarStochastic <: ScalarVariate
type ArrayStochastic{N} <: ArrayVariate{N}
typealias AbstractStochastic Union{ScalarStochastic, ArrayStochastic}
Fields¶
value: values of typeFloat64forScalarStochasticnodes andArray{Float64}forArrayStochasticnodes that represent samples from a target distribution.symbol::Symbol: identifying symbol for the node.monitor::Vector{Int}: indices identifying elements of thevaluefield to include in monitored MCMC sampler output.eval::Function: function for updating thedistrfield for the node.sources::Vector{Symbol}: other nodes upon whom the distributional specification for this one depends.targets::Vector{Symbol}:Dependentnodes that depend on this one. Elements oftargetsare topologically sorted so that a given node in the vector is conditionally independent of subsequent nodes, given the previous ones.distr: distributional specification of typeUnivariateDistributionforScalarStochasticnodes andDistributionStructforArrayStochasticnodes.
Distribution Structures¶
The DistributionStruct alias defines the types of distribution structures supported for AbstractStochastic nodes. Single Distribution types from the Distributions section, arrays of UnivariateDistribution, and arrays of MultivariateDistribution objects are supported. When a MultivariateDistribution array is specified for a stochastic node, the node is assumed to be one dimension bigger than the array, with the last dimension containing values from the distributions stored in the previous dimensions. Such arrays may contain distributions of different lengths. Model specification syntax for all three types of distribution structures can be seen in the Birats Example.
typealias DistributionStruct Union{Distribution,
Array{UnivariateDistribution},
Array{MultivariateDistribution}}
Constructors¶
-
Stochastic(f::Function, monitor::Union{Bool, Vector{Int}}=true)¶ -
Stochastic(d::Integer, f::Function, monitor::Union{Bool, Vector{Int}}=true) Construct a
Stochasticobject that defines a stochastic model node.Arguments
d: number of dimensions for array nodes.f: function whose untyped arguments are the other model nodes upon which this one depends. The function may contain any valid julia expression or code block. It will be saved in theevalfield of the constructed stochastic node and should return aDistributionStructobject to be stored in the node’sdistrfield.monitor: boolean indicating whether all elements are monitored, or vector of element-wise indices of elements to monitor.
Value
Returns anArrayStochasticif the dimension argumentdis specified, and aScalarStochasticif not.Example
See the Model Specification section of the tutorial.
Initialization¶
-
setinits!(s::Stochastic, m::Model, x=nothing) Set initial values for a stochastic node.
Arguments
s: stochastic node to which to assign initial values.m: model containing the node.x: values to assign to the node.
Value
Returns the node with its assigned initial values.
Node Operations¶
-
logpdf(s::AbstractStochastic, transform::Bool=false) -
logpdf(s::AbstractStochastic, x, transform::Bool=false) Evaluate the log-density function for a stochastic node.
Arguments
s: stochastic node for which to evaluate the log-density.x: value, of the same type and shape as the node value, at which to perform the evaluation. If not specified, the node value is used.transform: whether the evaluation is on the link-transformed scale.
Value
The resulting numeric value of the log-density.
-
rand(s::AbstractStochastic)¶ Draw a sample from the distributional specification on a stochastic node.
Arguments
s: stochastic node from which to generate a random sample.
Value
Returns the sampled value(s).
-
unlist(s::AbstractStochastic, transform::Bool=false) -
unlist(s::AbstractStochastic, x::Real, transform::Bool=false) -
unlist(s::AbstractStochastic, x::AbstractArray, transform::Bool=false) -
relist(s::AbstractStochastic, x::AbstractArray, transform::Bool=false) Extract (unlist) stochastic node values to a vector, or re-assemble (relist) values into a format that can be put into a node. These methods are used internally to extract the unique and sampled values of stochastic nodes. They are used, for instance, to extract only the unique, upper-triangular portions of (symmetric) covariance matrices and only the sampled values of
Array{MultivariateDistribution}specifications whose distributions may be of different lengths.Arguments
s: stochastic node for which to unlist or relist values.x: values to be listed. If not specified, the node values are used.transform: whether to apply a link transformation, or its inverse, to map values in a constrained distributional support to an unconstrained space. Supports for continuous, univariate distributions and positive-definite matrix distributions (Wishart or inverse-Wishart) are transformed as follows:- Lower and upper bounded: scaled and shifted to the unit interval and logit-transformed.
- Lower bounded: shifted to zero and log-transformed.
- Upper bounded: scaled by -1, shifted to zero, and log-transformed.
- Positive-definite matrix: compute the (upper-triangular) Cholesky decomposition, and return it with the diagonal elements log-transformed.
Value
Returns the extractedxvalues as a vector or the re-assembled values in the same shape as the specified node.
-
update!(s::AbstractStochastic, m::Model) Update the values of a stochastic node according to its relationship with others in a model.
Arguments
s: stochastic node to update.m: model containing the node.
Value
Returns the node with its values updated.