Distributions¶
Given in this section are distributions, as provided by the Distributions [1] and Mamba packages, supported for the specification of Stochastic nodes. Truncated versions of continuous univariate distributions are also supported.
Univariate Distributions¶
Distributions Package Univariate Types¶
The following univariate types from the Distributions package are supported.
Arcsine Cosine Hypergeometric NegativeBinomial Rayleigh
Bernoulli DiscreteUniform InverseGamma NoncentralBeta Skellam
Beta Edgeworth InverseGaussian NoncentralChisq TDist
BetaPrime Erlang KSDist NoncentralF TriangularDist
Binomial Exponential KSOneSided NoncentralT Uniform
Categorical FDist Laplace Normal Weibull
Cauchy Gamma Levy NormalCanon
Chi Geometric Logistic Pareto
Chisq Gumbel LogNormal Poisson
Flat Distribution¶
A Flat distribution is supplied with the degenerate probability density function:
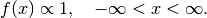
Flat() # Flat distribution
User-Defined Univariate Distributions¶
New known, unknown, or unnormalized univariate distributions can be created and added to Mamba as subtypes of the Distributions package ContinuousUnivariateDistribution or DiscreteUnivariateDistribution types. Mamba requires only a partial implementation of the method functions described in the full instructions for creating univariate distributions. The specific workflow is given below.
Create a
quoteblock for the new distribution. Assign the block a variable name, sayextensions, preceded by the@everywheremacro to ensure compatibility when julia is run in multi-processor mode.The Distributions package contains types and method definitions for new distributions. Load the package and import the package’s methods (indicated below) to be extended.
Declare a new distribution subtype, say
D, within the block. Create a constructor for the subtype that accepts un-typed arguments and explicitly converts them in the constructor body to the proper types for the fields ofD. Implementing the constructor in this way ensures that it will be callable with the MambaStochasticandLogicaltypes.Extend/define the following Distributions package methods for the new distribution
D.Test the subtype.
Add the
quoteblock (new distribution) to Mamba with the following calls.using Mamba @everywhere eval(Mamba, extensions)
Below is a univariate example based on the linear regression model in the Tutorial.
## Define a new univariate Distribution type for Mamba.
## The definition must be placed within an unevaluated quote block.
@everywhere extensions = quote
## Load needed packages and import methods to be extended
using Distributions
import Distributions: minimum, maximum, logpdf
## Type declaration
type NewUnivarDist <: ContinuousUnivariateDistribution
mu::Float64
sigma::Float64
## Constructor
function NewUnivarDist(mu, sigma)
new(convert(Float64, mu), convert(Float64, sigma))
end
end
## The following method functions must be implemented
## Minimum and maximum support values
minimum(d::NewUnivarDist) = -Inf
maximum(d::NewUnivarDist) = Inf
## Normalized or unnormalized log-density value
function logpdf(d::NewUnivarDist, x::Real)
-log(d.sigma) - 0.5 * ((x - d.mu) / d.sigma)^2
end
end
## Test the extensions in a temporary module (optional)
module Testing end
eval(Testing, extensions)
d = Testing.NewUnivarDist(0.0, 1.0)
Testing.minimum(d)
Testing.maximum(d)
Testing.insupport(d, 2.0)
Testing.logpdf(d, 2.0)
## Add the extensions to Mamba
using Mamba
@everywhere eval(Mamba, extensions)
## Implement a Mamba model using the new distribution
model = Model(
y = Stochastic(1,
@modelexpr(mu, s2,
begin
sigma = sqrt(s2)
Distribution[NewUnivarDist(mu[i], sigma) for i in 1:length(mu)]
end
),
false
),
mu = Logical(1,
@modelexpr(xmat, beta,
xmat * beta
),
false
),
beta = Stochastic(1,
:(MvNormal(2, sqrt(1000)))
),
s2 = Stochastic(
:(InverseGamma(0.001, 0.001))
)
)
## Sampling Scheme
scheme = [NUTS([:beta]),
Slice([:s2], [3.0])]
## Sampling Scheme Assignment
setsamplers!(model, scheme)
## Data
line = (Symbol => Any)[
:x => [1, 2, 3, 4, 5],
:y => [1, 3, 3, 3, 5]
]
line[:xmat] = [ones(5) line[:x]]
## Initial Values
inits = Dict{Symbol,Any}[
[:y => line[:y],
:beta => rand(Normal(0, 1), 2),
:s2 => rand(Gamma(1, 1))]
for i in 1:3]
## MCMC Simulation
sim = mcmc(model, line, inits, 10000, burnin=250, thin=2, chains=3)
describe(sim)
Multivariate Distributions¶
Distributions Package Multivariate Types¶
The following multivariate types from the Distributions package are supported.
Dirichlet Multinomial MvNormal MvNormalCannon
MvTDist VonMisesFisher
Block-Diagonal Multivariate Normal Distribution¶
A Block-Diagonal Multivariate Normal distribution is supplied with the probability density function:
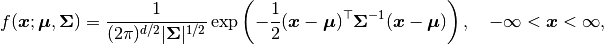
where
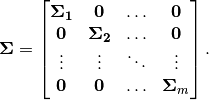
BDiagNormal(mu, C) # multivariate normal with mean vector mu and block-
# diagonal covariance matrix Sigma such that
# length(mu) = dim(Sigma), and Sigma_1 = ... = Sigma_m = C
# for a matrix C or Sigma_1 = C[1], ..., Sigma_m = C[m]
# for a vector of matrices C.
User-Defined Multivariate Distributions¶
New known, unknown, or unnormalized multivariate distributions can be created and added to Mamba as subtypes of the Distributions package ContinuousMultivariateDistribution or DiscreteMultivariateDistribution types. Mamba requires only a partial implementation of the method functions described in the full instructions for creating multivariate distributions. The specific workflow is given below.
Create a
quoteblock for the new distribution. Assign the block a variable name, sayextensions, preceded by the@everywheremacro to ensure compatibility when julia is run in multi-processor mode.The Distributions package contains types and method definitions for new distributions. Load the package and import the package’s methods (indicated below) to be extended.
Declare a new distribution subtype, say
D, within the block. Create a constructor for the subtype that accepts un-typed arguments and explicitly converts them in the constructor body to the proper types for the fields ofD. Implementing the constructor in this way ensures that it will be callable with the MambaStochasticandLogicaltypes.Extend/define the following Distributions package methods for the new distribution
D.Test the subtype.
Add the
quoteblock (new distribution) to Mamba with the following calls.using Mamba @everywhere eval(Mamba, extensions)
Below is a multivariate example based on the linear regression model in the Tutorial.
## Define a new multivariate Distribution type for Mamba.
## The definition must be placed within an unevaluated quote block.
@everywhere extensions = quote
## Load needed packages and import methods to be extended
using Distributions
import Distributions: length, insupport, _logpdf
## Type declaration
type NewMultivarDist <: ContinuousMultivariateDistribution
mu::Vector{Float64}
sigma::Float64
## Constructor
function NewMultivarDist(mu, sigma)
new(convert(Vector{Float64}, mu), convert(Float64, sigma))
end
end
## The following method functions must be implemented
## Dimension of the distribution
length(d::NewMultivarDist) = length(d.mu)
## Logicals indicating whether elements of x are in the support
function insupport{T<:Real}(d::NewMultivarDist, x::Vector{T})
length(d) == length(x) && all(isfinite(x))
end
## Normalized or unnormalized log-density value
function _logpdf{T<:Real}(d::NewMultivarDist, x::Vector{T})
-length(x) * log(d.sigma) - 0.5 * sumabs2(x - d.mu) / d.sigma^2
end
end
## Test the extensions in a temporary module (optional)
module Testing end
eval(Testing, extensions)
d = Testing.NewMultivarDist([0.0, 0.0], 1.0)
Testing.insupport(d, [2.0, 3.0])
Testing.logpdf(d, [2.0, 3.0])
## Add the extensions to Mamba
using Mamba
@everywhere eval(Mamba, extensions)
## Implement a Mamba model using the new distribution
model = Model(
y = Stochastic(1,
@modelexpr(mu, s2,
NewMultivarDist(mu, sqrt(s2))
),
false
),
mu = Logical(1,
@modelexpr(xmat, beta,
xmat * beta
),
false
),
beta = Stochastic(1,
:(MvNormal(2, sqrt(1000)))
),
s2 = Stochastic(
:(InverseGamma(0.001, 0.001))
)
)
## Sampling Scheme
scheme = [NUTS([:beta]),
Slice([:s2], [3.0])]
## Sampling Scheme Assignment
setsamplers!(model, scheme)
## Data
line = (Symbol => Any)[
:x => [1, 2, 3, 4, 5],
:y => [1, 3, 3, 3, 5]
]
line[:xmat] = [ones(5) line[:x]]
## Initial Values
inits = Dict{Symbol,Any}[
[:y => line[:y],
:beta => rand(Normal(0, 1), 2),
:s2 => rand(Gamma(1, 1))]
for i in 1:3]
## MCMC Simulation
sim = mcmc(model, line, inits, 10000, burnin=250, thin=2, chains=3)
describe(sim)
Matrix-Variate Distributions¶
Distributions Package Matrix-Variate Types¶
The following matrix-variate types from the Distributions package are supported.
InverseWishart Wishart