Model¶
The Model type is designed to store the set of all model nodes, including parameter set 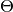 as denoted in the Mamba Gibbs sampling scheme. In particular, it stores
as denoted in the Mamba Gibbs sampling scheme. In particular, it stores Dependent type objects in its nodes dictionary field. Valid models are ones whose nodes form directed acyclic graphs (DAGs). Sampling functions 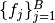 are saved as
are saved as Sampler objects in the vector of field samplers. Vector elements 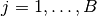 correspond to sampling blocks
correspond to sampling blocks 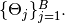
Declaration¶
type Model
Fields¶
nodes::Dict{Symbol, Any}: all input, logical, and stochastic model nodes.samplers::Vector{Sampler}: sampling functions for updating blocks of stochastic nodes.states::Vector{ModelState}: states of chains at the end of an MCMC run in a possible series of runs, whereModelStatehas fieldsvalue::Vector{Float64}andtune::Vector{Any}to store the last values of sampled nodes and block-sampler tuning parameters, respectively.iter::Int: current MCMC draw from the target distribution.burnin::Int: number of initial draws to discard as a burn-in sequence to allow for convergence.hasinputs::Bool: whether values have been assigned to input nodes.hasinits::Bool: whether initial values have been assigned to stochastic nodes.
Constructor¶
-
Model(; iter::Integer=0, burnin::Integer=0, samplers::Vector{Sampler}=Sampler[], nodes...)¶ Construct a
Modelobject that defines a model for MCMC simulation.Arguments
iter: current iteration of the MCMC simulation.burnin: number of initial draws to be discarded as a burn-in sequence to allow for convergence.samplers: block-specific sampling functions.nodes...: arbitrary number of user-specified arguments defining logical and stochastic nodes in the model. Argument values must beLogicalorStochastictype objects. Their names in the model will be taken from the argument names.
Value
Returns aModeltype object.Example
See the Model Specification section of the tutorial.
MCMC Engine¶
-
mcmc(m::Model, inputs::Dict{Symbol}, inits::Vector{Dict{Symbol, Any}}, iters::Integer; burnin::Integer=0, thin::Integer=1, chains::Integer=1, verbose::Bool=true)¶ -
mcmc(mc::ModelChains, iters::Integer; verbose::Bool=true) Simulate MCMC draws for a specified model.
Arguments
m: specified model.mc: chains from a previous call tomcmcfor which to simulate additional draws.inputs: values for input model nodes. Dictionary keys and values should be given for each input node.inits: dictionaries that contain initial values for stochastic model nodes. Dictionary keys and values should be given for each stochastic node. Consecutive runs of the simulator will iterate through the vector’s dictionary elements.iters: number of draws to generate for each simulation run.burnin: numer of initial draws to discard as a burn-in sequence to allow for convergence.thin: step-size between draws to output.chains: number of simulation runs to perform.verbose: whether to print sampler progress at the console.
Value
AModelChainstype object of simulated draws.Example
See the MCMC Simulation section of the tutorial.
Indexing¶
-
getindex(m::Model, nodekey::Symbol)¶ Returns a model node identified by its symbol. The syntax
m[nodekey]is converted togetindex(m, nodekey).Arguments
m: model containing the node to get.nodekey: node to get.
Value
The specified node.
-
keys(m::Model)¶ -
keys(m::Model, ntype::Symbol, at...) Extract the symbols (keys) for all existing nodes or for nodes of a specified type.
Arguments
m: model containing the nodes of interest.ntype: type of nodes to return. Options are:all: all input, logical, and stochastic model nodes.:assigned: nodes that have been assigned values.:block: stochastic nodes being updated by the sampling block(s)at::Integer=0(default: all blocks).:dependent: logical and stochastic (dependent) nodes in topologically sorted order.:independentor:input: input (independent) nodes.:logical: logical nodes.:monitor: stochastic nodes being monitored in MCMC sampler output.:output: stochastic nodes upon which no other stochastic nodes depend.:source: nodes upon which the nodeat::Symbolor vector of nodesat::Vector{Symbol}depends.:stochastic: stochastic nodes.:target: topologically sorted nodes that depend on the sampling block(s)at::Integer=0(default: all blocks), nodeat::Symbol, or vector of nodesat::Vector{Symbol}.
at...: additional positional arguments to be passed to thentypeoptions, as described above.
Value
A vector of node symbols.
Display¶
-
draw(m::Model; filename::AbstractString="")¶ Draw a GraphViz DOT-formatted graph representation of model nodes and their relationships.
Arguments
m: model for which to construct a graph.filename: external file to which to save the resulting graph, or an empty string to draw to standard output (default). If a supplied external file name does not include a dot (.), the file extension.dotwill be appended automatically.
Value
The model drawn to an external file or standard output. Stochastic, logical, and input nodes will be represented by ellipses, diamonds, and rectangles, respectively. Nodes that are unmonitored in MCMC simulations will be gray-colored.Example
See the Directed Acyclic Graphs section of the tutorial.
-
graph(m::Model)¶ Construct a graph representation of model nodes and their relationships.
Arguments
m: model for which to construct a graph.
Value
Returns aModelGraphtype object with fieldgraphcontaining aDiGraphrepresentation of indices, as defined in the LightGraphs package, to a vector of node symbols in fieldkeys.
-
graph2dot(m::Model)¶ Draw a GraphViz DOT-formatted graph representation of model nodes and their relationships.
Arguments
m: model for which to construct a graph.
Value
A character string representation of the graph suitable for in-line processing. Stochastic, logical, and input nodes will be represented by ellipses, diamonds, and rectangles, respectively. Nodes that are unmonitored in MCMC simulations will be gray-colored.Example
See the Directed Acyclic Graphs section of the tutorial.
-
show(m::Model)¶ Write a text representation of the model, nodes, and attributes to the current output stream.
-
showall(m::Model)¶ Write a verbose text representation of the model, nodes, and attributes to the current output stream.
Initialization¶
-
setinits!(m::Model, inits::Dict{Symbol, Any})¶ Set the initial values of stochastic model nodes.
Arguments
m: model with nodes to be initialized.inits: initial values for stochastic model nodes. Dictionary keys and values should be given for each stochastic node.
Value
Returns the model with stochastic nodes initialized and theiterfield set equal to 0.Example
See the Development and Testing section of the tutorial.
-
setinputs!(m::Model, inputs::Dict{Symbol, Any})¶ Set the values of input model nodes.
Arguments
m: model with input nodes to be assigned.inputs: values for input model nodes. Dictionary keys and values should be given for each input node.
Value
Returns the model with values assigned to input nodes.Example
See the Development and Testing section of the tutorial.
-
setsamplers!(m::Model, samplers::Vector{T<:Sampler})¶ Set the block-samplers for stochastic model nodes.
Arguments
m: model with stochastic nodes to be sampled.samplers: block-specific samplers.
Values:
Returns the model updated with the block-samplers.Example
See the Model Specification and MCMC Simulation sections of the tutorial.
Parameter Block Operations¶
-
gettune(m::Model, block::Integer=0)¶ Get block-sampler tuning parameters.
Arguments
m: model with block-samplers.block: block for which to get the tuning parameters (default: all blocks).
Value
AVector{Any}of all block-specific tuning parameters ifblock=0, and turning parameters for the specified block otherwise.
-
gradlogpdf(m::Model, block::Integer=0, transform::Bool=false; dtype::Symbol=:forward)¶ -
gradlogpdf(m::Model, x::AbstractVector{T<:Real}, block::Integer=0, transform::Bool=false; dtype::Symbol=:forward) -
gradlogpdf!(m::Model, x::AbstractVector{T<:Real}, block::Integer=0, transform::Bool=false; dtype::Symbol=:forward)¶ Compute the gradient of log-densities for stochastic nodes.
Arguments
m: model containing the stochastic nodes for which to compute the gradient.block: sampling block of stochastic nodes for which to compute the gradient (default: all stochastic nodes).x: value (possibly different than the current one) at which to compute the gradient.transform: whether to compute the gradient of block parameters on the link–transformed scale.dtype: type of differentiation for gradient calculations. Options are:central: central differencing.:forward: forward differencing.
Value
The resulting gradient vector. Methodgradlogpdf!()additionally updates modelmwith supplied valuesx.Note
Numerical approximation of derivatives by central and forward differencing is performed with the Calculus package [97].
-
logpdf(m::Model, block::Integer=0, transform::Bool=false)¶ -
logpdf(m::Model, nodekeys::Vector{Symbol}, transform::Bool=false) -
logpdf(m::Model, x::AbstractArray{T<:Real}, block::Integer=0, transform::Bool=false) -
logpdf!(m::Model, x::AbstractArray{T<:Real}, block::Integer=0, transform::Bool=false)¶ Compute the sum of log-densities for stochastic nodes.
Arguments
m: model containing the stochastic nodes for which to evaluate log-densities.block: sampling block of stochastic nodes over which to sum densities (default: all stochastic nodes).nodekeys: nodes over which to sum densities.x: value (possibly different than the current one) at which to evaluate densities.transform: whether to evaluate evaluate log-densities of block parameters on the link–transformed scale.
Value
The resulting numeric value of summed log-densities. Methodlogpdf!()additionally updates modelmwith supplied valuesx.
-
sample!(m::Model, block::Integer=0)¶ Generate one MCMC sample of values for a specified model.
Argument:
m: model specification.block: block for which to sample values (default: all blocks).
Value
Returns the model updated with the MCMC sample and, in the case ofblock=0, theiterfield incremented by 1.Example
See the Development and Testing section of the tutorial.
-
unlist(m::Model, block::Integer=0, transform::Bool=false)¶ -
unlist(m::Model, nodekeys::Vector{Symbol}, transform::Bool=false) -
relist(m::Model, x::AbstractArray{T<:Real}, block::Integer=0, transform::Bool=false)¶ -
relist(m::Model, x::AbstractArray{T<:Real}, nodekeys::Vector{Symbol}, transform::Bool=false) -
relist!(m::Model, x::AbstractArray{T<:Real}, block::Integer=0, transform::Bool=false)¶ -
relist!(m::Model, x::AbstractArray{T<:Real}, nodekey::Symbol, transform::Bool=false) Convert (unlist) sets of logical and/or stochastic node values to vectors, or reverse (relist) the process.
Arguments
m: model containing nodes to be unlisted or relisted.block: sampling block of nodes to be listed (default: all blocks).nodekey/nodekeys: node(s) to be listed.x: values to re-list.transform: whether to apply a link transformation in the conversion.
Value
Theunlistmethods return vectors of concatenated node values,relistreturn dictionaries of symbol keys and values for the specified nodes, andrelist!return their model argument with values copied to the nodes.
-
update!(m::Model, block::Integer=0)¶ -
update!(m::Model, nodekeys::Vector{Symbol}) Update values of logical and stochastic model node according to their relationship with others in a model.
Arguments
m: mode with nodes to be updated.block: sampling block of nodes to be updated (default: all blocks).nodekeys: nodes to be updated in the given order.
Value
Returns the model with updated nodes.